Home
Dommage à l’ADN, réparation et maintenance du génome
L’exposition des organismes aux agents environnementaux tels que les rayons ultraviolets du soleil, les rayons ionisants et de nombreux produits génotoxiques, cause des dommages à l’ADN. S’ils ne sont pas réparés, ces dommages induisent la formation de mutations qui pourraient éventuellement amener au développement de cancers. Ainsi, la stabilité du génome est importante dans la prévention de l’oncogenèse.
Nous nous intéressons, tout particulièrement, aux mécanismes de réparation de l’ADN endommagé. De plus, dans les cellules l’ADN est condensé dans une structure appelée chromatine. Les recherches de notre laboratoire ont comme but de comprendre comment l’ADN est réparé au niveau de la chromatine. Nous pensons qu’une meilleure compréhension des mécanismes de réparation de l’ADN pourra aider à définir leur rôle dans la prévention du cancer.
D’un autre côté, plus de la moitié des médicaments utilisés en chimiothérapie provoquent des dommages au niveau de l’ADN. Ces dommages bloquent la transcription et la réplication de l’ADN et donc inhibent la croissance cellulaire. Les dommages causés par la plupart des agents utilisés en chimiothérapie sont cependant aussi réparés par les mécanismes de réparation de l’ADN. L’étude de ces mécanismes, au niveau de la chromatine, pourra aider à synthétiser des agents chimiothérapeutiques plus efficaces, par exemple ciblant des régions de la chromatine qui sont plus difficiles à réparer et perturberait alors la réplication de l’ADN. Nous nous intéressons donc aux modes d’actions des médicaments chimiothérapeutiques qui endommagent l’ADN.
DNA damage, DNA repair and maintenance of the genome
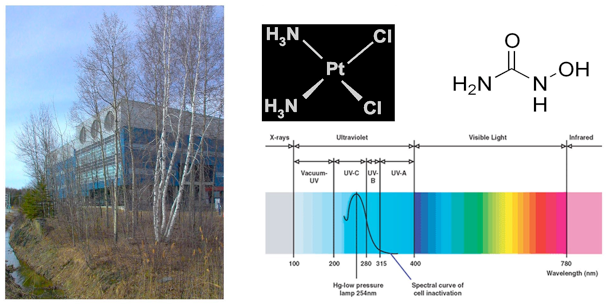
Environmental agents such as the ultraviolet (UV) component of sunlight, ionizing radiation, numerous genotoxic chemicals and pollutants cause DNA damage. If not repaired, DNA damage can lead to mutations and increased risk for cancer. For example, UV light induces two major types of damage: cyclobutane pyrimidine dimers and (6-4) photoproducts, both of which are associated with skin cancer.
The genome is organized into nuclear sub-domains, which create microenvironments favoring distinct chromatin structures and functions (e.g., highly repetitive sequences, centromeres, telomeres, inactive genes, RNA polymerase-III, -II and -I transcribed genes). Correlations have been drawn between gene silencing and its proximity to a heterochromatic compartment. For instance, yeast telomeres are clustered at the nuclear periphery, are formed onto an altered chromatin structure that shares features with heterochromatin, and silences adjacent genes. At the other end of the scale are ribosomal genes, which are transcribed at a very high rate by RNA polymerase I (~60% of total transcription), have a loose chromatin structure and are clustered in the nucleolus. Thus, we propose that the kinetics of DNA repair vary among the nuclear sub-domains. To better understand the mechanisms of DNA repair and how cells maintain the integrity of the genome, we analyze DNA repair in different chromosomal contexts.
There are 4 categories of DNA repair: Recombination-, Mismatch-, Base excision- and Nucleotide excision (NER)- repair. The repair pathways have been conserved through evolution and are present from bacteria to human. In general, these repair pathways function independently since they are rather specialized for different groups of DNA lesions. However, under certain circumstances extensive cross-talk can occur between repair pathways. In the laboratory we study NER, which is performed by a large multi-enzymatic complex and removes numerous types of lesions from the DNA, including: bulky adducts caused by chemicals, inter- or intra-strand cross-links and UV photoproducts. NER in yeast presents most of the hallmarks of NER in human cells. The strength of the yeast system relies on both genetic and biochemical approaches, which combined make a powerful tool to study intricate processes such as DNA repair in chromatin. Currently, we study NER in RNA polymerase I, -II and -III transcribed genes (in their active and inactive chromatin states), and NER of the specialized chromatin structure at the chromosomes ends.
On the other hand, nearly half of the drugs used in chemotherapy make DNA lesions. In fact, DNA damage induced by these compounds block DNA transcription and replication, thereby arresting cell proliferation. Therefore, a better understanding of DNA repair in living cells is of paramount importance to health science, and is the objective of the ongoing research in our laboratory.